Surprising insight into Legionnaires' disease
 July 2019. In order to control cellular processes and thwart the immune system, the bacterium Legionella pneumophilia, the cause of the notorious Legionnaires' disease, releases hundreds of enzymes. Biochemists at Goethe University have now elucidated important details of the interaction of bacterial effectors. They discovered how the regulatory enzyme SidJ keeps other dangerous virulence factors in check.
July 2019. In order to control cellular processes and thwart the immune system, the bacterium Legionella pneumophilia, the cause of the notorious Legionnaires' disease, releases hundreds of enzymes. Biochemists at Goethe University have now elucidated important details of the interaction of bacterial effectors. They discovered how the regulatory enzyme SidJ keeps other dangerous virulence factors in check.
The incidence of Legionnaires' disease has increased in the past two decades. The natural habitat of Legionella is freshwater biotopes, where they mainly reproduce in amoebae. In addition, Legionella can also colonize water tanks or pipes and spread, for example, via poorly maintained air-conditioning systems. Contaminated aerosols are released into the air and trigger the infection. The pathogen causes also pneumonia, which is often fatal in elderly patients or individuals with a weak immune system.
What makes Legionella so dangerous is its ability to multiply in phagocytes of the human immune system by secreting virulence factors. Some of these effectors - the enzymes of what is known as the SidE family - are so toxic that without tight control they would instantly kill their host cells. However, since Legionella needs the host cell in order to multiply, it has developed a sophisticated mechanism for the precise control of SidE enzyme activity. Details of this process are now reported by scientists at Goethe University and from Grenoble in the journal Nature.
 The scientists have shown that another protein, the regulator SidJ, which is
also released by Legionella, works as an antidote to SidE enzymes.
SidJ thus ensures accurate control of SidE activity. The SidJ regulator is
a glutamylase, i.e. it has a rare enzymatic activity that allows amino acid
glutamate molecules to be linked together to form chains. In this case,
SidJ attacks the central glutamate of SidE and inhibits its activity. So far,
little is known about glutamylases and the scientists were surprised when
they discovered that it is precisely this type of enzyme which is important
for the coordinated interaction of the virulence factors of Legionella.
The scientists have shown that another protein, the regulator SidJ, which is
also released by Legionella, works as an antidote to SidE enzymes.
SidJ thus ensures accurate control of SidE activity. The SidJ regulator is
a glutamylase, i.e. it has a rare enzymatic activity that allows amino acid
glutamate molecules to be linked together to form chains. In this case,
SidJ attacks the central glutamate of SidE and inhibits its activity. So far,
little is known about glutamylases and the scientists were surprised when
they discovered that it is precisely this type of enzyme which is important
for the coordinated interaction of the virulence factors of Legionella.
"This is a typical example of how completely unpredictable results drive research. Such discoveries are what make science such a fascinating and exciting profession", says Ivan Dikic from the Institute of Biochemistry II and the Buchmann Institute for Molecular Life Sciences at Goethe University Frankfurt. "We can only gain a molecular understanding of the complex world of bacterial infections by working in interdisciplinary teams and combining methods from modern biochemistry and proteomics with cell and structural biology techniques."
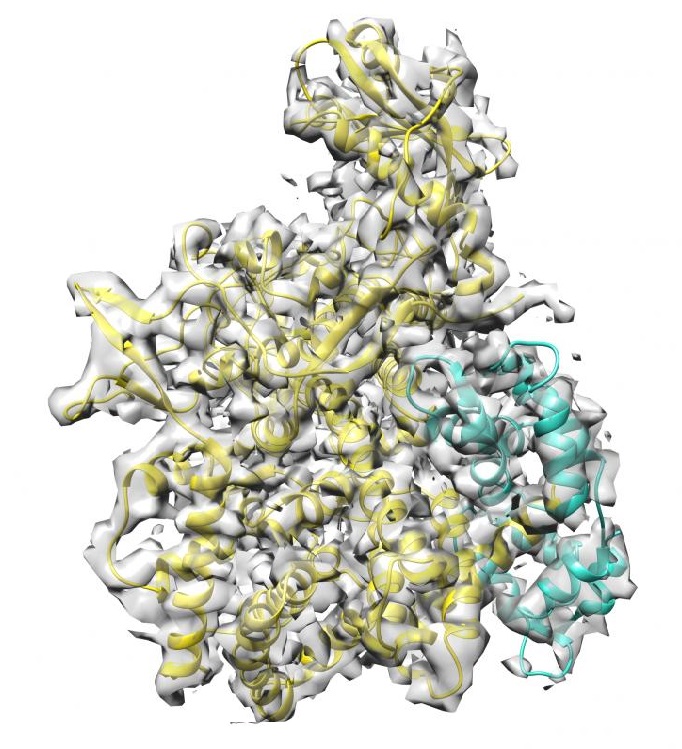
The researchers also revealed how SidJ is activated in host cells: It requires the calcium-binding protein calmodulin found in mammalian cells. Cryo-electron microscopy played an important role in explaining the structure of the calmodulin-SidJ complex. "Glutamylation as a protein modification is understudied. Our finding that Legionella pneumophilia uses exactly this mechanism to sustain the infection certainly argues for more research in this field. For example, the extent to which Legionella utilizes this modification to regulate other cellular processes is completely unclear", explains Sagar Bhogaraju, who led the microscopic examinations at the European Molecular Biology Laboratory (EMBL) in Grenoble.
This so far unknown mechanism opens up new possibilities for research to inhibit the spread of Legionella in the host organism. "We're currently working on eliminating SidJ selectively by developing inhibitors for the glutamylase domain. In addition to the use of antibiotics, they could prevent the spread of Legionella pneumophilia in phagocytes", explains Dikic.
Publication:
Sagar Bhogaraju, Florian Bonn, Rukmini Mukherjee, Michael Adams, Moritz M. Pfleiderer, Wojciech P. Galej , Vigor Matkovic, Sissy Kalayil, Donghyuk Shin1, Ivan Dikic: Inhibition of SidE ubiquitin ligases through SidJ/Calmodulin catalyzed glutamylation, in Nature 22. Juli 2019 DOI: 10.1038/s41586-019-1440-8
(Link)
Contact:
Ivan Dikic, Buchmann Institute for Molecular Life Sciences and Institute of Biochemistry II, Goethe University Frankfurt, Frankfurt/Main, Germany, dikic(at)biochem2.uni-frankfurt.de.